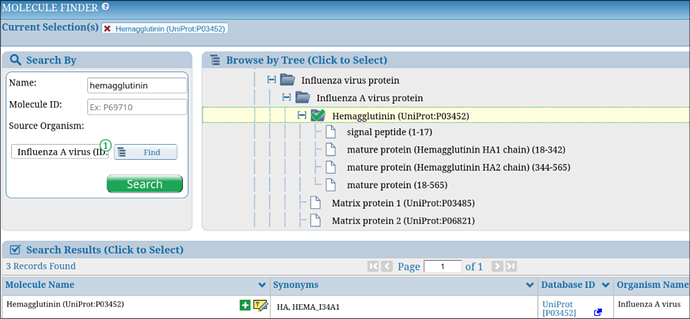
The Molecule Finder is a tool designed to help users explore and select source antigens of epitopes within the Immune Epitope Database (IEDB). You can access it directly from the IEDB homepage or on the results page in the “Epitope Source” box, located next to the “Antigen” field. The Molecule Finder provides access to the protein sources of natural peptidic epitopes and also non-peptidic structures containing smaller non-peptidic epitopes. The Molecule Finder offers two ways to locate molecules: browsing a tree structure that organizes non-peptidic and protein source data, or using the search bars on the left to query by name, molecule ID, or source organism. For example, searching for “Influenza A” in the Source Organism field and “hemagglutinin” in the name field quickly retrieves relevant results, as shown in Figure 1.
Figure 1. Example of searching Influenza A hemagglutinin in the Molecule Finder.
Non-peptidic molecules are sourced from the Chemical Entities of Biological Interest (ChEBI) database, each having a ChEBI ID, which is displayed as the molecule’s IRI in data exports. Additionally, synonyms for non-peptidic structures are imported from ChEBI and can be found in the Synonyms column in data exports.
The protein branch relies on mappings from curated source antigens—proteins identified in research papers—to a “parent protein” within a selected proteome. This mapping simplifies queries; for instance, with Influenza A hemagglutinin, numerous strain-specific antigens exist in the literature, but users can focus on a top-level hemagglutinin protein. Proteomes are sourced from UniProt and chosen at the species level based on specific criteria outlined below.
Proteomes and Protein Mapping
Proteomes for the Molecule Finder are derived from UniProt and classified into five types:
- Representative: Selected to represent a species.
- Reference: Chosen for well-studied model organisms.
- Non-redundant: A proteome distinct from others with high similarity.
- Other: Generated computationally from genetic data but not fully annotated.
- Orphans: Not considered an “official” proteome, but are taxonomic mappings to individual protein entries in the UniProt Knowledgebase.
For each species, we select the highest-ranked proteome. In cases of ties, we prioritize the proteome with the most epitope matches to ensure relevance to immunological data. These proteomes serve as the foundation for mapping source antigens.
Source antigens, which are proteins (or their isoforms) identified in curated research papers along with their associated epitopes, are aligned to the selected proteome using either BLAST or MMseqs2. For species with many antigens, we opt for MMseqs2, which searches approximately 50 times faster than BLAST. The process identifies the top-matching protein hit and its associated gene. We then use PEPMatch, a peptide search tool, to search for epitopes across all isoforms of that gene, assigning each epitope to its best match. This ensures that users can query epitope data efficiently, linking to their parent proteins within the proteome.
Additional Features and Search Options
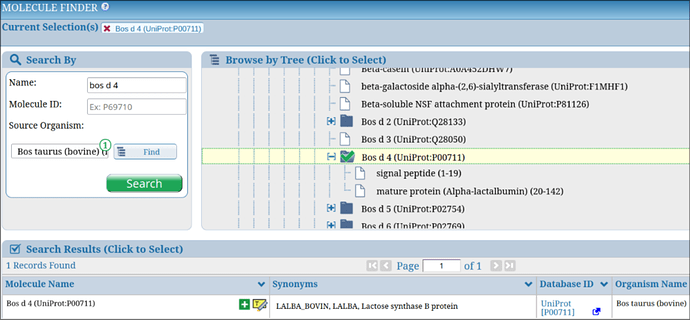
The Molecule Finder supports specialized immunological nomenclature to enhance usability. For allergens, it incorporates official International Union of Immunological Societies (IUIS) terms, such as “Bos d 4” for alpha-lactalbumin from cow’s milk (Figure 2), allowing allergen researchers to search using familiar identifiers. These IUIS terms are mapped to their corresponding proteins within the selected proteomes, ensuring seamless integration with epitope data.
For vertebrate species, the Finder also includes organizational nodes for immune receptor proteins: Major Histocompatibility Complex (MHC), B-cell receptors (BCR) or immunoglobulin (Ig), and T-cell receptors (TCR). These nodes group related proteins under a single category, making it easier to explore epitope data associated with immune recognition molecules. For example, selecting the MHC node for a species like Homo sapiens reveals all curated MHC-related antigens and their epitopes.
The Molecule Finder also supports the display of protein fragments, such as signal peptides, transit peptides, or mature protein segments, which are derived from UniProt data. These fragments are listed as child nodes under their parent protein in the molecule tree, allowing users to explore epitope data associated with specific protein regions. For example, as shown in Figure 2, searching for “Bos d 4” reveals its signal peptide and mature protein as distinct child entries, enabling precise queries for epitopes linked to these fragments.
Figure 2. Example of searching for the allergen “Bos d 4” in the Molecule Finder, showing its placement in the protein tree along with its fragmented entries.