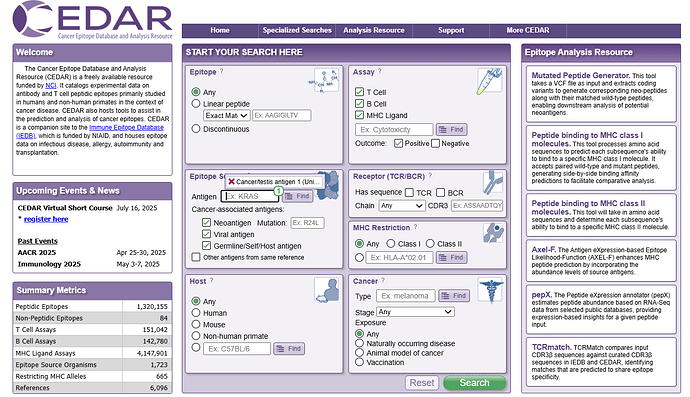
Users can use the ‘Epitope Source’ panel to query CEDAR for epitopes encoded by a specific gene by entering it into the text field. When the user starts to enter text into the field, matching proteins will show up in a drop-down menu. In the example query in Figure 1, the antigen ‘NY-ESO-1’ was selected. The ‘Molecule Finder’ can be accessed by clicking the ‘Find’ button next to the text field. Here, users can search for a gene/protein of interest by entering a synonym or the UniProt ID and the source organism.
Figure 1. The Epitope Source panel
Additionally, users can select specific cancer-associated antigen subtypes as the epitope source, including ‘Neoantigen’, ‘Viral antigen’, and ‘Germline/Self/Host antigen’. We defined these three broader categories of cancer-associated antigens that can be clearly distinguished and are mutually exclusive. Antigens that are not cancer-associated but were reported together with cancer-associated antigens in a cancer-related study can also be included in the query by selecting ‘Other antigens from same reference’. The default selection on the CEDAR homepage excludes those antigens and only queries all cancer-associated antigens. In the example query in Figure 1, the checkboxes ‘Neoantigen’ and ‘Viral antigen’ were deselected to include only germline epitopes.