Introduction to the CEDAR Query API
Welcome to the new CEDAR Query API! We have now made it possible to programmatically query the CEDAR using multiple endpoints, enabling users to complete most queries available from the CEDAR home page and work with the data directly in their preferred environment. We hope this help article provides additional context on the new CEDAR API.
If you have any questions or feedback, please contact us at cedar@lji.org.
What is the CEDAR API?
The CEDAR API is built upon a PostgREST platform that allows for transparent access to the Postgres tables on the backend. Each table can be queried through individual endpoints, described in this interactive Swagger documentation.
What endpoints are available?
The CEDAR API provides two types of endpoints: search and export. The search endpoints contain multiple fields with information that facilitate programmatic identification and/or filter the data of interest, while the export endpoints match the structure and naming conventions of the custom exports on the CEDAR website.
Core search endpoints
- epitope_search
- antigen_search
- tcell_search (assays)
- bcell_search (assays)
- mhc_search (assays)
- tcr_search (receptors)
- bcr_search (receptors)
- reference_search
Export endpoints
- epitope_export
- tcell_export (assays)
- bcell_export (assays)
- mhc_export (assays)
- tcr_export (assays)
- bcr_export (assays)
- reference_export
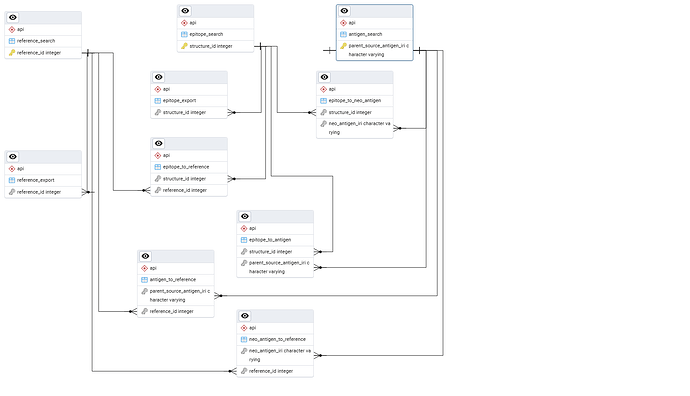
As shown in the Entity Relationship Diagram (ERD), these endpoints are interconnected by different fields (more ERDs can be found in this github repository). We recommend first querying the relevant search table and then using resource embeddings to retrieve the needed data from its associated export table. An example of resource embeddings usage is provided at the end of this document. The entire list of available fields is described in the interactive Swagger documentation.
Supporting endpoints
An additional endpoint, ‘curie_map’, links CURIE prefixes (e.g., PMID) to their full IRIs. Further details on CURIEs and IRIs can be found below.
Finally, the ‘api_metrics’ endpoint provides record counts and build dates for the core endpoints.
How to query the CEDAR API?
As the CEDAR API is based on PostgREST, queries must be performed using its rich and expressive query syntax, described here in detail. Detailed API walkthroughs are available in our CEDAR API github repository as Python notebooks.
The most basic example of querying for the first 10 epitopes is provided here, using the ‘curl’ command.
curl "https://cedar-api.iedb.org/epitope_search?limit=10" | jq
[
{
"structure_id": 68100,
"structure_iri": "IEDB_EPITOPE:68100",
"structure_descriptions": [
"VDYLKEKTKDQDL"
],
"curated_source_antigens": null,
…
Only the first part of the response is shown, as the full response includes many fields. Note the ‘pipe’ to ‘jq’, used to format the data nicely for display. By default, results are returned in JSON format. If CSV (comma-separated values) format is preferred, an additional header has to be provided in the GET query. The same query would become:
curl "https://cedar-api.iedb.org/epitope_search?limit=10" -H "accept: text/csv"
The output is not shown, as there are too many fields to display. However, we can limit the output to a subset of fields with the ‘select’ parameter. For example, if we only want the ‘structure_id’ and ‘linear_sequence’ of the first 10 epitopes and we want it returned in CSV format, the query becomes:
curl "https://cedar-api.iedb.org/epitope_search?limit=10&select=structure_id,linear_sequence" -H "accept: text/csv"
structure_id,linear_sequence
585066,KPIAPCGAPSK
585140,KPSIDLMSKW
123811,VEFDNEVQ
585954,LHQLGREALAI
585955,LHSLSLVLTL
123813,VGAVQGPV
586048,LLRTGYTQAHSL
586065,LMPKRVLEI
586068,LMYASSKDAIK
586069,LNAEKNSLL
As already mentioned, the search endpoints contain particular fields that facilitate the programmatic identification and/or filtering of relevant data. One example of these fields is the host_organism_iri_search, which contains the complete taxonomic lineage of the host organism, including all parent taxa. This enables queries at any taxonomic rank (e.g., kingdom, order, species) by matching against the corresponding identifier in the hierarchy. The taxonomies are expressed as IRIs (Internationalized Resource Identifiers), which provide stable links to external resources, such as the NCBI Taxonomy. Let’s include this field to inspect its structure:
curl "https://cedar-api.iedb.org/epitope_search?limit=10&select=structure_id,linear_sequence,host_organism_iri_search" -H "accept: text/csv"
structure_id,linear_sequence,host_organism_iri_search
585066,KPIAPCGAPSK,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
585140,KPSIDLMSKW,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
123811,VEFDNEVQ,"{NCBITaxon:314147,NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606,NCBITaxon:9986,taxon:10000206}"
585954,LHQLGREALAI,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
585955,LHSLSLVLTL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
123813,VGAVQGPV,"{NCBITaxon:314147,NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606,NCBITaxon:9986,taxon:10000206}"
586048,LLRTGYTQAHSL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586065,LMPKRVLEI,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586068,LMYASSKDAIK,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586069,LNAEKNSLL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
Now, if we want to retrieve epitopes only tested in humans as hosts, the query would be:
curl "https://cedar-api.iedb.org/epitope_search?limit=10&select=structure_id,linear_sequence,host_organism_iri_search&host_organism_iri_search.ov.{NCBITaxon:9606}" -H "accept: text/csv"
structure_id,linear_sequence,host_organism_iri_search
585066,KPIAPCGAPSK,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
585140,KPSIDLMSKW,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
123811,VEFDNEVQ,"{NCBITaxon:314147,NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606,NCBITaxon:9986,taxon:10000206}"
585954,LHQLGREALAI,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
585955,LHSLSLVLTL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
123813,VGAVQGPV,"{NCBITaxon:314147,NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606,NCBITaxon:9986,taxon:10000206}"
586048,LLRTGYTQAHSL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586065,LMPKRVLEI,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586068,LMYASSKDAIK,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
586069,LNAEKNSLL,"{NCBITaxon:314295,NCBITaxon:40674,NCBITaxon:9443,NCBITaxon:9606}"
All the fields in the search endpoints denoted with the suffix ‘_search’ have this ‘tree like’ structure, and are connected to different ontologies, such as ONTIE, GeneOntology, DiseaseOntology, and OntoBee, among others. Further details on CEDAR Ontologies can be found in this article.
Other examples of search endpoint fields supporting complex queries are the neoantigen_bool, viral_antigen_bool, and germline_antigen_bool fields, which behave as the checkboxes on the Epitope Source panel of the CEDAR home page. For example, if we want to retrieve neoantigens and germline antigens, the query would be:
curl "https://cedar-api.iedb.org/epitope_search?limit=10&select=structure_id,linear_sequence&host_organism_iri_search.ov.{NCBITaxon:9606}&or=(neoantigen_bool.eq.1,germline_antigen_bool.eq.1)" -H "accept: text/csv"
structure_id,linear_sequence
119671,WLNGKEYK
132802,ESTLHLVLRLR
128259,AKKAGAAKAKKPAGAA
136144,AVQPLLLGRIIASYD
136687,TLKAGGILNRFSKDI
136733,VTSLKGLWTLRAFGR
137402,FIDSYICQV
2256759,GPLHKCDISNSTEAG
2256915,NTILNTMSTIYSTGK
2256885,MLRLGKSEPWTLALE
What are IDs, IRIs, and CURIEs?
Several types of identifiers are used throughout the database to track unique records. First, there are internal integer record identifiers denoted with the suffix ‘_id’, e.g., ‘reference_id’. These are generally in the first field of each table. As they are internal to the CEDAR, they cannot be linked directly to other resources.
Many of the tables in the database also have fields that end in ‘_iri’, e.g., ‘reference_iri’. These are identifiers that resolve uniquely and unambiguously to records both within and outside of the CEDAR. The Internationalized Resource Identifier (IRI) specification includes Uniform Resource Locators (URLs), which we use as globally unique identifiers, e.g., https://cedar.iedb.org/reference/1043674. A shortened version of an IRI, called a CURIE, can be constructed by replacing a portion of the IRI with a common prefix. The above IRI can be represented in CURIE format as ‘IEDB_REFERENCE:1043674.
By querying against the ‘curie_map’ endpoint, it is possible to find prefixes for converting between the two representations, e.g.: https://cedar-api.iedb.org/curie_map?limit=3
curl "https://cedar-api.iedb.org/curie_map?limit=5&select=curie_prefix,iri_replace" -H "accept: text/csv"
curie_prefix,iri_replace
PMID,https://www.ncbi.nlm.nih.gov/pubmed/?term=
IEDB_REFERENCE,https://www.iedb.org/reference/
IEDB_SUBMISSION,https://www.iedb.org/submission/
IEDB_EPITOPE,https://www.iedb.org/epitope/
IEDB_COMPLEX,https://www.iedb.org/3dViewer.php?complex=
IRIs for Antigens
Only for the ‘antigen_id’ field in the ‘antigen_search’ table, an IRI is used rather than an internal integer record identifier.
Troubleshooting and FAQs
Common issues and idiosyncrasies
Curated vs Parent Terms
There are some terms called “curated”, such as curated_source_antigens, while other similar terms use the phrase “parent”, such as parent_source_antigen_names.
“Curated” refers to the precise source protein isoform that matches exactly what an author referred to as the source of a peptide epitope in a specific publication. The curated_source_antigen will 100% BLAST match to the epitope sequence.
“Parent” refers to a reference protein that serves as a representative protein for all the isoforms to which an epitope was linked in different studies. The parent groups these isoforms together, but it will not always be an exact BLAST match to the epitope sequence.
Similarly, the source_organism_name (shown nested under curated_source_antigens) reflects the precise source organism strain that matches exactly what an author referred to as the source of an epitope in a specific publication. While the parent_source_antigen_source_org_name is the species-level organism name that groups all strains that might ever have been associated with that same epitope across all publications.
This help desk article goes into further detail: Epitope Source.
Results Page Limit & Default Page Size
By default, the CEDAR API has a maximum page size of 10,000 records. In practice, this means that queries that result in more than 10,000 results will be divided into pages, and only the first 10,000 records will be returned by the initial query.
NOTE: If a query requires paging, it is critical to also provide the ‘order’ parameter to determine how the rows are sorted. If it is not provided, rows will be returned in a random order, and pages will be inconsistent between queries.
The API will always return a count of the records matching the query, as well as the number of pages of results. This information is embedded in the ‘content-range’ response header, e.g.:
curl -I "https://cedar-api.iedb.org/antigen_search"
HTTP/1.1 200 OK
Date: Thu, 04 Sep 2025 23:12:03 GMT
Server: postgrest/9.0.1
Content-Range: 0-9999/*
Content-Location: /antigen_search
Content-Type: application/json; charset=utf-8
The server returned the first 10,000 records (indexed as 0-9999). The trailing ‘/*’ indicates that there are more records matching the query but the total number has not been calculated. To get an exact count of matching records, you must provide the header ‘Prefer: count=exact’ in your GET request.
curl -I "https://cedar-api.iedb.org/antigen_search" -H 'Prefer: count=exact'
HTTP/1.1 206 Partial Content
Date: Thu, 04 Sep 2025 23:13:33 GMT
Server: postgrest/9.0.1
Content-Range: 0-9999/66340
Content-Location: /antigen_search
Content-Type: application/json; charset=utf-8
Now we can see that there are 66,340 matching records, which would correspond to 8 pages of results. Note that adding this header can be detrimental to query performance. To retrieve the last page of the results, we add the ‘offset’ parameter to the query:
curl -I "https://cedar-api.iedb.org/antigen_search?offset=66340&order=parent_source_antigen_id" -H 'Prefer: count=exact'
HTTP/1.1 206 Partial Content
Date: Thu, 04 Sep 2025 23:18:01 GMT
Server: postgrest/9.0.1
Content-Range: */66340
Content-Location: /antigen_search?offset=66340&order=parent_source_antigen_id
Content-Type: application/json; charset=utf-8
If an offset is defined that is higher than the number of matching records, an empty result will be returned.
Error messages
Large Query Error - “Cannot enlarge string buffer containing…out of memory”
This message is the result of a large amount of information being returned. Many of the tables in the database contain fields that are information-dense, which can cause buffering issues on the PostGres backend. If a user receives this message, the recommended workflow is to:
- Try adding a ‘limit’ parameter to your query to fetch the first N (e.g., 100) records
- Simultaneously, add the request header ‘Prefer: count=exact’ so you are aware of the total number of records matching the query
- Update the query to filter rows (on field values) or unnecessary columns (using ‘select’) and/or continue to page through the results by adding an ‘offset’ parameter
FAQs
– Can I save API links from the CEDAR?
Not yet - this feature is currently in development. You will soon be able to retrieve the relevant API links on the ‘Results’ page using the ‘Export’ function.
– Can I query for epitope sequence similarity to BLAST matches at 90%, 80% and 70%?
Unfortunately, the blast match options can not be replicated via the CEDAR API due to limitations on how it’s calculated. The CEDAR API can only be used for ‘Exact’ and ‘Substring’ queries. For example:
- Exact match (you will want to do an equals search eq.): https://cedar-api.iedb.org/epitope_search?linear_sequence=eq.SIINFEKL
- Substring match (you will want to do a like search with a * wildcard on either side of the search term): https://cedar-api.iedb.org/epitope_search?linear_sequence=like.*SIINFEKL*
– Can I search for multiple epitopes from the CEDAR web interface?
The web interface currently only allows searching one sequence at a time. However, the CEDAR API allows users to simultaneously query for multiple epitope sequences via these endpoints.
– Is there a way to access the number of subjects or response frequency for specific assays through the CEDAR API?
Yes, however, these values are not part of the assay search endpoints. To access the additional fields available in the export endpoints while still running standard searches in the search endpoints, one can rely on PostgREST Resource Embedding. This feature allows retrieval of information from another linked endpoint within the current endpoint. For example, in the case of bcell_search:
https://cedar-api.iedb.org/bcell_search?select=*,bcell_export(assay__number_of_subjects_tested,assay__number_of_subjects_positive,assay__response_frequency_)&bcell_id=eq.1384867
As shown in this example, the bcell_search query uses the assay ID to link to the bcell_export information, pulling the specified fields from that endpoint into the existing API call.
Contact information
Contact us via email at cedar@lji.org to provide your feedback.